In accordance to the Earth Wellness Corporation, every single yr there are an estimated 1 billion situations of influenza, among 3-5 million critical scenarios and up to 650,000 influenza-related respiratory fatalities globally. Seasonal flu vaccines need to be reformulated each individual yr to match the predominantly circulating strains. When the vaccine matches the predominant pressure, it is very productive having said that, when it does not match, it might give very little security.
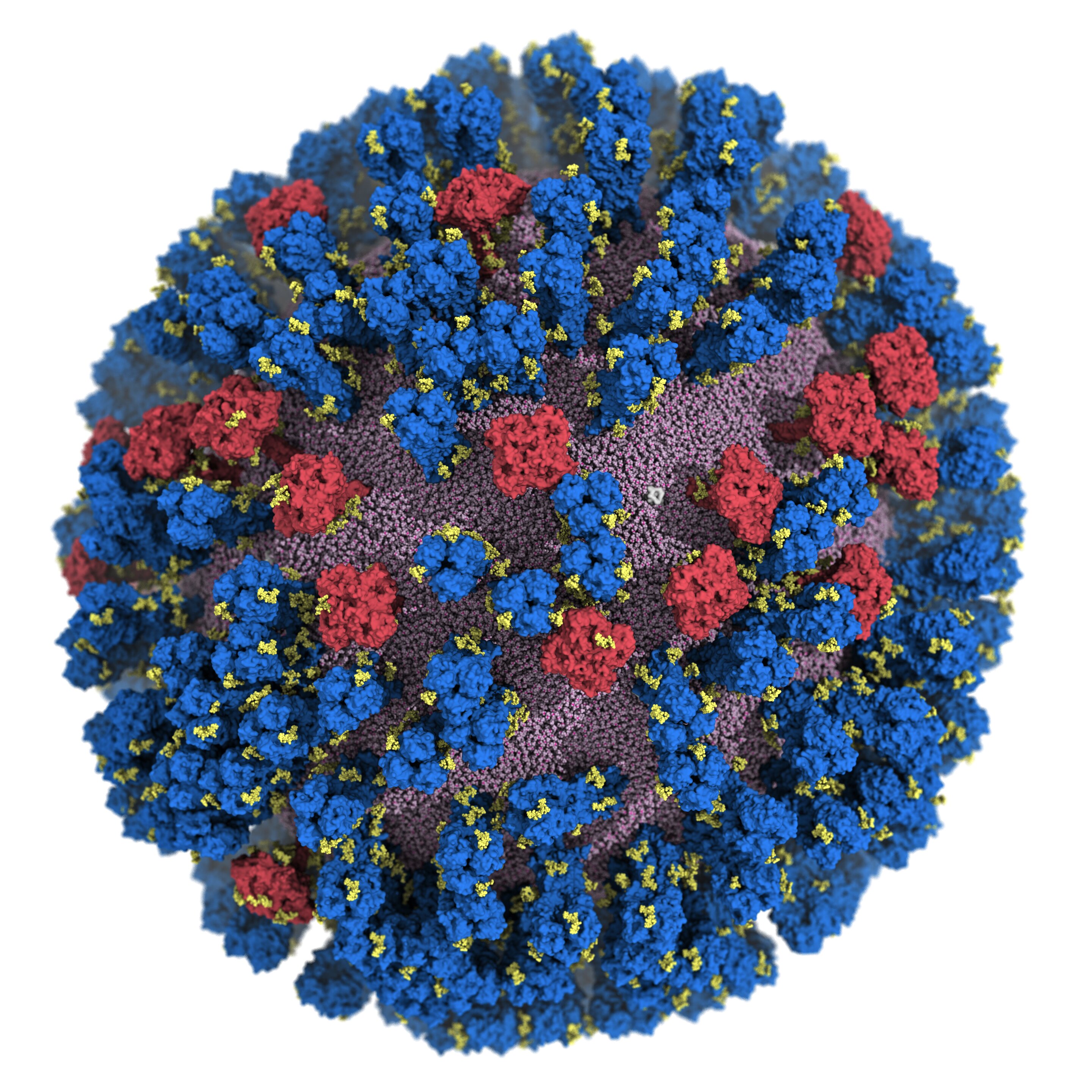
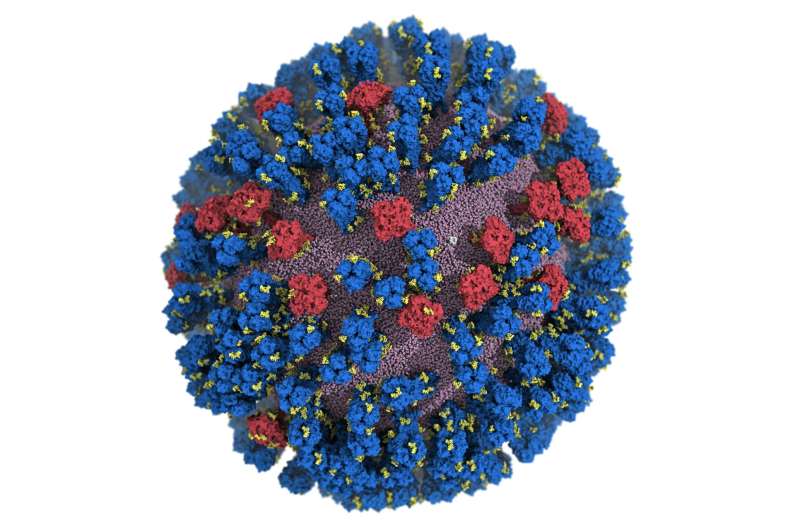
The most important targets of the flu vaccine are two surface glycoproteins, hemagglutinin (HA) and neuraminidase (NA). When the HA protein allows the virus bind to the host mobile, the NA protein functions like scissors to minimize the HA absent from the mobile membrane allowing the virus to replicate. While the properties of the two glycoproteins have been examined earlier, a entire comprehending of their movement does not exist.
For the initial time, scientists at the University of California San Diego have developed an atomic-amount personal computer product of the H1N1 virus that reveals new vulnerabilities by means of glycoprotein “respiratory” and “tilting” movements. This function, released in ACS Central Science, suggests probable techniques for the structure of foreseeable future vaccines and antivirals from influenza.
“When we very first observed how dynamic these glycoproteins have been, the substantial degree of respiratory and tilting, we truly questioned if there was some thing erroneous with our simulations,” stated Distinguished Professor of Chemistry and Biochemistry Rommie Amaro, who is the principal investigator on the task. “At the time we understood our versions have been correct, we understood the great potential this discovery held. This investigate could be used to create methods of maintaining the protein locked open up so that it would be consistently accessible to antibodies.”
Customarily, flu vaccines have focused the head of the HA protein based on even now visuals that showed the protein in a restricted formation with little motion. Amaro’s model showed the dynamic nature of the HA protein and uncovered a respiratory movement that exposed a formerly unfamiliar web-site of immune response, identified as an epitope.
This discovery complemented previous work from just one of the paper’s co-authors, Ian A. Wilson, Hansen Professor of Structural Biology at The Scripps Research Institute, who experienced found an antibody that was broadly neutralizing—in other text, not pressure-specific—and sure to a element of the protein that appeared unexposed. This suggested that the glycoproteins had been far more dynamic than earlier considered, allowing for the antibody an possibility to attach. Simulating the respiration motion of the HA protein founded a link.
NA proteins also confirmed motion at the atomic degree with a head-tilting motion. This delivered a key insight to co-authors Julia Lederhofer and Masaru Kanekiyo at the Nationwide Institute of Allergy and Infectious Health conditions. When they looked at convalescent plasma—that is, plasma from individuals recovering from the flu—they discovered antibodies specially concentrating on what is identified as the “dark side” of NA underneath the head.
Without having observing the movement of NA proteins, it was not crystal clear how the antibodies had been accessing the epitope. The simulations Amaro’s lab established showed an remarkable selection of motion that gave insight into how the epitope was uncovered for antibody binding.
The H1N1 simulation Amaro’s workforce developed consists of an huge total of detail—160 million atoms well worth. A simulation of this measurement and complexity can only run on a few choose devices in the world. For this function, the Amaro lab utilized Titan at Oak Ridge National Lab, previously one particular of the most significant and fastest computer systems in the entire world.
Amaro is producing the info obtainable to other scientists who can uncover even a lot more about how the influenza virus moves, grows and evolves. “We’re mainly fascinated in HA and NA, but there are other proteins, the M2 ion channel, membrane interactions, glycans, so several other opportunities,” Amaro mentioned.
“This also paves the way for other teams to apply equivalent procedures to other viruses. We have modeled SARS-CoV-2 in the earlier and now H1N1, but there are other flu variants, MERS, RSV, HIV—this is just the beginning.”
Extra information:
Lorenzo Casalino et al, Respiration and Tilting: Mesoscale Simulations Illuminate Influenza Glycoprotein Vulnerabilities, ACS Central Science (2022). DOI: 10.1021/acscentsci.2c00981
Quotation:
Laptop model of influenza virus exhibits common vaccine assure (2023, January 25)
retrieved 28 January 2023
from https://phys.org/information/2023-01-influenza-virus-common-vaccine.html
This doc is subject matter to copyright. Aside from any truthful dealing for the purpose of personal research or research, no
portion may possibly be reproduced devoid of the published permission. The articles is provided for facts purposes only.